quantms.io tools¶
quantms.io tools provides a standardized set of commands to generate different files for your project. It is mainly used to consolidate the data of each file and generate a standardized file representation.
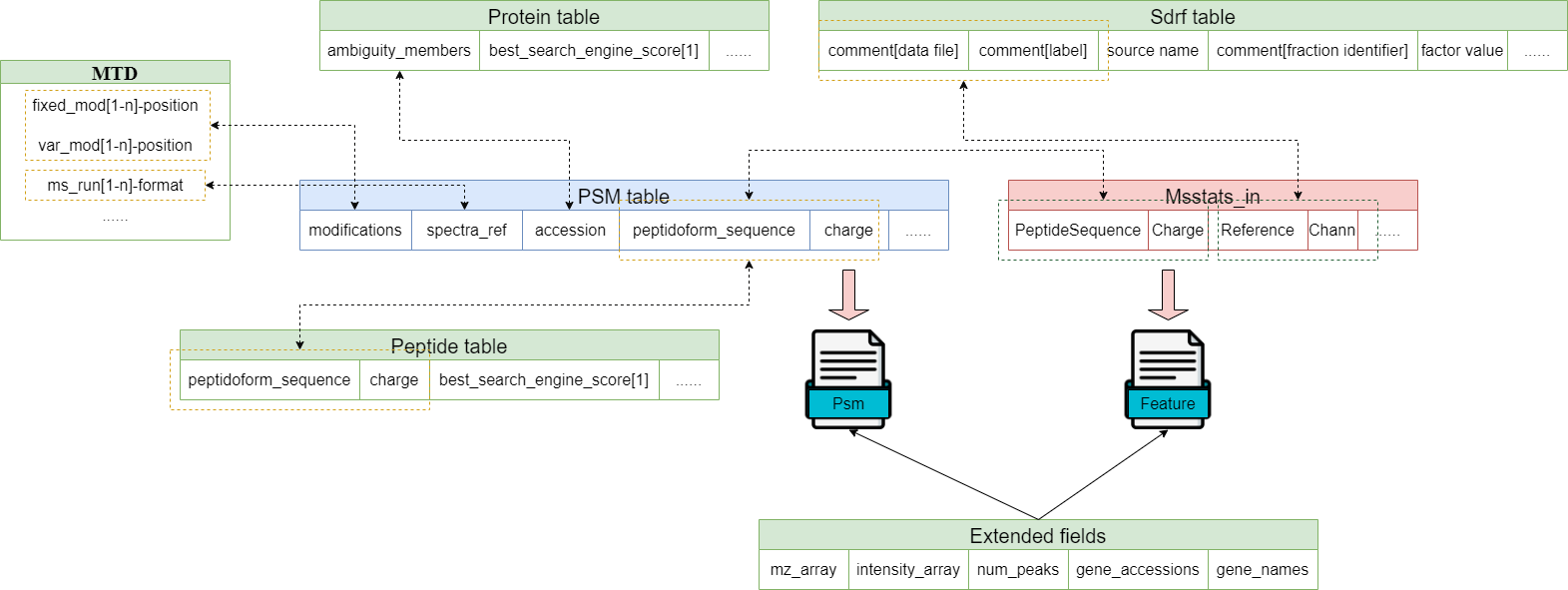
You can generate separate files or complete project files depending on your needs.A completed project contains the following files:
project.json-> contains descriptive information about the entire project.
Example:
{
"project_accession": "PXD014414",
"project_title": "",
"project_sample_description": "",
"project_data_description": "",
"project_pubmed_id": 32265444,
"organisms": [
"Homo sapiens"
],
"organism_parts": [
"mammary gland",
"adjacent normal tissue"
],
"diseases": [
"metaplastic breast carcinomas",
"Triple-negative breast cancer",
"Normal",
"not applicable"
],
"cell_lines": [
"not applicable"
],
"instruments": [
"Orbitrap Fusion"
],
"enzymes": [
"Trypsin"
],
"experiment_type": [
"Triple-negative breast cancer",
"Wisp3",
"Tandem mass tag (tmt) labeling",
"Ccn6",
"Metaplastic breast carcinoma",
"Precision therapy",
"Lc-ms/ms shotgun proteomics"
],
"acquisition_properties": [
{"proteomics data acquisition method": "TMT"},
{"proteomics data acquisition method": "Data-dependent acquisition"},
{"dissociation method": "HCD"},
{"precursor mass tolerance": "20 ppm"},
{"fragment mass tolerance": "0.6 Da"}
],
"quantms_files": [
{"feature_file": "PXD014414-943a8f02-0527-4528-b1a3-b96de99ebe75.featrue.parquet"},
{"sdrf_file": "PXD014414-f05eca35-9381-40d8-a7da-2fe57745afaf.sdrf.tsv"},
{"psm_file": "PXD014414-f4fb88f6-0a45-451d-a8a6-b6d58fb83670.psm.parquet"},
{"differential_file": "PXD014414-3026e5d5-fb0e-45e9-a4f0-c97d86536716.differential.tsv"}
],
"quantms_version": "1.1.1",
"comments": []
}
absolute_expression.tsvordifferential_expression.tsv
The differential expression format by quantms is based on the MSstats output.
Example:
protein |
label |
log 2fc |
se |
d f |
pv al ue |
adj.p value |
i ss ue |
|---|---|---|---|---|---|---|---|
LV86 1_HUMAN |
normal-squamous cell carcinoma |
0 .60 |
0. 87 |
8 |
0. 51 |
0.62 |
NA |
The absolute expression format by quantms contains IBAQ message.
Example:
protein |
sample_accession |
condition |
ibaq |
ribaq |
|---|---|---|---|---|
LV861_HUMAN |
Sample-1 |
heart |
1234.1 |
12.34 |
feature.parquet
The feature.parquet cover detail on peptide level.
Example:
sequence |
protein_accessions |
protein_start_positions |
protein_end_positions |
protein_global_qvalue |
unique |
modifications |
retention_time |
charge |
exp_mass_to_charge |
calc_mass_to_charge |
peptidoform |
posterior_error_probability |
global_qvalue |
is_decoy |
intensity |
spectral_count |
sample_accession |
condition |
fraction |
biological_replicate |
fragment_ion |
isotope_label_type |
run |
channel |
id_scores |
reference_file_name |
best_psm_reference_file_name |
best_psm_scan_number |
mz_array |
intensity_array |
num_peaks |
gene_accessions |
gene_names |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
ASPDWGYDDK |
[‘sp|CONTAMINANT_P00915|CONTAMINANT_CAH1_HUMAN’,’sp|P00915|CAH1_HUMAN’] |
[1 2] |
[10 11] |
0.001882796 |
0 |
[‘0-UNIMOD:1’ ‘10-UNIMOD:737’] |
7522.223146 |
2 |
712.831298 |
712.8302134 |
[Acetyl]-ASPDWGYDDK[TMT6plex] |
4.97E-05 |
0 |
0 |
454585.3 |
1 |
PXD014414-Sample-10 |
Norm |
1 |
10 |
None |
L |
1_1_1 |
TMT131 |
[“‘OpenMS:Best PSM Score’:0.0”,’Best PSM PEP:4.96872e-05’] |
UM_F_50cm_2019_0414 |
UM_F_50cm_2019_0430 |
53434 |
psm.parquet
psm.parquet store details on PSM level including spectrum mz/intensity for specific use-cases such as AI/ML training.
Example:
sequence |
protein_accessions |
protein_start_positions |
protein_end_positions |
protein_global_qvalue |
unique |
modifications |
retention_time |
charge |
exp_mass_to_charge |
calc_mass_to_charge |
peptidoform |
posterior_error_probability |
global_qvalue |
is_decoy |
id_scores |
consensus_support |
reference_file_name |
scan_number |
mz_array |
intensity_array |
num_peaks |
gene_accessions |
gene_names |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
SSPGHR |
[‘sp|P29692|EF1D_HUMAN’] |
[118] |
[123] |
0.001882796 |
1 |
[‘1-UNIMOD:737’] |
1258.2 |
2 |
435.2432855 |
435.2431809 |
S[TMT6plex]SPGHR |
0.35875 |
0 |
[“‘OpenMS:Target-decoy PSM q-value’: 0.040626999360205”,’Posterior error probability: 0.35875’] |
UM_F_50cm_2019_0428 |
2193 |
sdrf.tsv
sdrf.tsv is a file used by quantMS to search the library.
Example:
source name |
characteristics[organism] |
characteristics[organism part] |
characteristics[developmental stage] |
characteristics[disease] |
characteristics[histologic subtype] |
characteristics[sex] |
characteristics[age] |
characteristics[cell type] |
characteristics[cell line] |
characteristics[biological replicate] |
characteristics[individual] |
Material Type |
assay name |
Technology Type |
comment[label] |
comment[data file] |
comment[file uri] |
comment[technical replicate] |
comment[fraction identifier] |
comment[cleavage agent details] |
comment[instrument] |
comment[modification parameters] |
comment[modification parameters] |
comment[modification parameters] |
comment[modification parameters] |
comment[modification parameters] |
comment[modification parameters] |
comment[dissociation method] |
comment[collision energy] |
comment[precursor mass tolerance] |
comment[fragment mass tolerance] |
factor value[disease] |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
PXD014414-Sample-1 |
Homo sapiens |
mammary gland |
adult |
metaplastic breast carcinomas |
Chondroid |
female |
43Y |
not applicable |
not applicable |
1 |
C1 |
tissue |
run 1 |
proteomic profiling by mass spectrometry |
TMT126 |
UM_F_50cm_2019_0414.raw |
ftp://ftp.pride.ebi.ac.uk/pride/data/archive/2020/04/PXD014414/UM_F_50cm_2019_0414.raw |
1 |
1 |
AC=MS:1001251;NT=Trypsin |
NT=Orbitrap Fusion;AC=MS:1002416 |
NT=Oxidation;MT=Variable;TA=M;AC=UNIMOD:35 |
NT=Acetyl;AC=UNIMOD:1;PP=Protein N-term;MT=variable |
NT=Carbamidomethyl;TA=C;MT=fixed;AC=UNIMOD:4 |
NT=TMT6plex;AC=UNIMOD:737;TA=K;MT=Fixed |
NT=TMT6plex;AC=UNIMOD:737;PP=Protein N-term;MT=Variable |
NT=TMT6plex;AC=UNIMOD:737;TA=S;MT=Variable |
NT=HCD;AC=PRIDE:0000590 |
55 NCE |
20 ppm |
0.6 Da |
metaplastic breast carcinomas |
If you want see a full example, please click here
Project converter tool¶
If your project comes from the PRIDE database,
you can use the pride accession to generate a project.json that contains
descriptive information about the entire project.
Or, customize a Project Accession to generate an entirely new project.
If you want to know more, please read Project file.
If your project is not from PRIDE, you can skip this step.
quantmsio_cli generate-pride-project-json
--project_accession PXD014414
--sdrf PXD014414.sdrf.tsv
--output_folder result
Optional parameter
--quantms_version Quantms version
--delete_existing Delete existing files in the output folder(default False)
DE converter tool¶
Differential expression file Store the differential express proteins between two contrasts, with the corresponding fold changes and p-values.It can be easily visualized using tools such as Volcano Plot and easily integrated with other omics data resources.
If you have generated project.json, you can use this parameter
--project_fileto add project information for DE files.If you want to know more, please read Differential expression format.
Example:
quantmsio_cli convert-de
--msstats_file PXD014414.sdrf_openms_design_msstats_in_comparisons.csv
--sdrf_file PXD014414.sdrf.tsv
--output_folder result
Optional parameter
--project_file Descriptive information from project.json(project json path)
--fdr_threshold FDR threshold to use to filter the results(default 0.05)
--output_prefix_file Prefix of the df expression file(like {prefix}-{uu.id}-{extension})
--delete_existing Delete existing files in the output folder(default True)
AE converter tool¶
The absolute expression format aims to visualize absolute expression (AE) results using iBAQ values and store the AE results of each protein on each sample.
If you have generated project.json, you can use this parameter
--project_fileto add project information for AE files.If you want to know ibaq, please read ibaqpy
If you want to know more, please read Absolute expression format.
Example:
quantmsio_cli convert-ae
--ibaq_file PXD004452-ibaq.csv
--sdrf_file PXD014414.sdrf.tsv
--output_folder result
Optional parameter
--project_file Descriptive information from project.json(project json path)
--output_prefix_file Prefix of the df expression file(like {prefix}-{uu.id}-{extension})
--delete_existing Delete existing files in the output folder(default True)
Feature converter tool¶
The Peptide table aims to cover detail on peptide level including peptide intensity. The most of content are from peptide part of mzTab. It store peptide intensity to perform down-stream analysis and integration.
If you want to know more, please read Peptide quantification features.
In some projects, mzTab files can be very large, so we provide both diskcache and no-diskcache versions of the tool.
You can choose the desired version according to your server configuration.
Example:
quantmsio_cli convert-feature
--sdrf_file PXD014414.sdrf.tsv
--msstats_file PXD014414.sdrf_openms_design_msstats_in.csv
--mztab_file PXD014414.sdrf_openms_design_openms.mzTab
--output_folder result
Optional parameter
--use_cache Whether to use diskcache instead of memory(default True)
--output_prefix_file The prefix of the result file(like {prefix}-{uu.id}-{extension})
--consensusxml_file The consensusXML file used to retrieve the mz/rt(default None)
Psm converter tool¶
The PSM table aims to cover detail on PSM level for AI/ML training and other use-cases. It store details on PSM level including spectrum mz/intensity for specific use-cases such as AI/ML training.
If you want to know more, please read PSM table format.
Example:
quantmsio_cli convert-psm
--mztab_file PXD014414.sdrf_openms_design_openms.mzTab
--output_folder result
Optional parameter
--use_cache Whether to use diskcache instead of memory(default True)
--output_prefix_file The prefix of the result file(like {prefix}-{uu.id}-{extension})
--verbose Output debug information(default True)
DiaNN convert¶
For DiaNN, the command supports generating feature.parquet and psm.parquet directly from diann_report files.
If you want to see
design_file, please click sdrf-pipelines
Example:
quantmsio_cli convert-diann
--report_path diann_report.tsv
--design_file PXD037682.sdrf_openms_design.tsv
--qvalue_threshold 0.05
--mzml_info_folder mzml
--sdrf_path PXD037682.sdrf.tsv
--output_folder result
--output_prefix_file PXD037682
Optional parameter
--duckdb_max_memory The maximum amount of memory allocated by the DuckDB engine (e.g 4GB)
--duckdb_threads The number of threads for the DuckDB engine (e.g 4)
--file_num The number of files being processed at the same time (default 100)
Inject some messages for DiaNN¶
For DiaNN, some field information is not available and needs to be filled with other commands.
bset-psm-scan-number
Example:
quantmsio_cli inject-bset-psm-scan-number
--diann_psm_path PXD010154-f75fbb29-4419-455f-a011-e4f776bcf73b.psm.parquet
--diann_feature_path PXD010154_map_protein_accession-88d63fca-3ae6-4eab-9262-6e7a68184432.feature.parquet
--output_path PXD010154.feature.parquet
start-and-end-pisition
Example:
quantmsio_cli inject-start-and-end-from-fasta
--parquet_path PXD010154_map_protein_accession-88d63fca-3ae6-4eab-9262-6e7a68184432.feature.parquet
--fasta_path Homo-sapiens-uniprot-reviewed-contaminants-decoy-202210.fasta
--label feature
--output_path PXD010154.feature.parquet
Compare psm.parquet¶
This tool is used to compare peptide information in result files obtained by different search engines.
--tagsor-tare used to specify the tags of the PSM table.
Example:
quantmsio_cli compare-set-psms
-p PXD014414-comet.parquet
-p PXD014414-sage.parquet
-p PXD014414-msgf.parquet
-t comet
-t sage
-t msgf
Generate spectra message¶
generate_spectra_message support psm and feature. It can be used directly for spectral clustering.
--labelcontains two options:psmandfeature.--partioncontains two options:chargeandreference_file_name.
Since the result file is too large, you can specify –-partition to split the result file.
Example:
quantmsio_cli map-spectrum-message-to-parquet
--parquet_path PXD014414-f4fb88f6-0a45-451d-a8a6-b6d58fb83670.psm.parquet
--mzml_directory mzmls
--output_path psm/PXD014414.parquet
--label psm
--file_num(default 10)
--partition charge
Generate gene message¶
generate_gene_message support psm and feature.
--labelcontains two options:psmandfeature.--map_parametercontains two options:map_protein_nameormap_protein_accession.
Example:
quantmsio_cli map-gene-msg-to-parquet
--parquet_path PXD000672-0beee055-ae78-4d97-b6ac-1f191e91bdd4.featrue.parquet
--fasta_path Homo-sapiens-uniprot-reviewed-contaminants-decoy-202210.fasta
--output_path PXD000672-gene.parquet
--label feature
--map_parameter map_protein_name
Optional parameter
--species species type(default human)
species
Common name |
Genus name |
|---|---|
human |
Homo sapiens |
mouse |
Mus musculus |
rat |
Rattus norvegicus |
fruitfly |
Drosophila melanogaster |
nematode |
Caenorhabditis elegans |
zebrafish |
Danio rerio |
thale-cress |
Arabidopsis thaliana |
frog |
Xenopus tropicalis |
pig |
Sus scrofa |
Map proteins accessions¶
get_unanimous_name support parquet and tsv. For parquet, map_parameter
have two option (map_protein_name or map_protein_accession), and the
label controls whether it is PSM or Feature.
parquet
--labelcontains two options:psmandfeature
Example:
quantmsio_cli labels convert-accession
--parquet_path PXD014414-f4fb88f6-0a45-451d-a8a6-b6d58fb83670.psm.parquet
--fasta Reference fasta database
--output_path psm/PXD014414.psm.parquet
--map_parameter map_protein_name
--label psm
tsv
Example:
quantmsio_cli labels get-unanimous-for-tsv
--path PXD014414-c2a52d63-ea64-4a64-b241-f819a3157b77.differential.tsv
--fasta Reference fasta database
--output_path psm/PXD014414.de.tsv
--map_parameter map_protein_name
Compare two parquet files¶
This tool is used to compare the feature.parquet file generated by two versions (diskcache or no-diskcache).
Example:
quantmsio_cli compare-parquet
--parquet_path_one res_lfq2_discache.parquet
--parquet_path_two res_lfq2_no_cache.parquet
--report_path report.txt
Generate report about files¶
This tool is used to generate report about all project.
Example:
quantmsio_cli generate-project-report
--project_folder PXD014414
Register file¶
This tool is used to register the file to project.json.
If your project comes from the PRIDE database, You can use this command to add file information for project.json.
The parameter
--categoryhas three options:feature_file,psm_file,differential_file,absolute_file.You can add the above file types.The parameter
--replace_existingis enable then we remove the old file and add this one. If not then we can have a list of files for a category.
Example:
quantmsio_cli attach-file
--project_file PXD014414/project.json
--attach_file PXD014414-943a8f02-0527-4528-b1a3-b96de99ebe75.featrue.parquet
--category feature_file
--replace_existing
Convert file to json¶
This tool is used to convert file to json.
parquet
--data_typecontains two options:psmandfeature
Example:
quantmsio_cli convert-parquet-json
--data_type feature
--parquet_path PXD014414-943a8f02-0527-4528-b1a3-b96de99ebe75.featrue.parquet
--json_path PXD014414.featrue.json
tsv
Example:
quantmsio_cli json convert-tsv-to-json
--file PXD010154-51b34353-227f-4d38-a181-6d42824de9f7.absolute.tsv
--json_path PXD010154.ae.json
sdrf
Example:
quantmsio_cli json convert-sdrf-to-json
--file MSV000079033-Blood-Plasma-iTRAQ.sdrf.tsv
--json_path MSV000079033.sdrf.json
Statistics¶
This tool is used for statistics. Example:
quantmsio_cli project-ae-statistics
--absolute_path PXD010154-51b34353-227f-4d38-a181-6d42824de9f7.absolute.tsv
--parquet_path PXD010154-51b34353-227f-4d38-a181-6d42824de9f7.featrue.parquet
--save_path PXD014414.statistic.txt
quantmsio_cli parquet-psm-statistics
--parquet_path PXD010154-51b34353-227f-4d38-a181-6d42824de9f7.psm.parquet
--save_path PXD014414.statistic.txt
Plots¶
This tool is used for visualization. - plot-psm-peptides
quantmsio_cli plot plot-psm-peptides
--psm_parquet_path PXD010154-51b34353-227f-4d38-a181-6d42824de9f7.psm.parquet
--sdrf_path PXD010154.sdrf.tsv
--save_path PXD014414_psm_peptides.svg
plot-ibaq-distribution
quantmsio_cli plot plot-ibaq-distribution
--ibaq_path PXD010154-51b34353-227f-4d38-a181-6d42824de9f7.ibaq.tsv
--select_column IbaqLog
--save_path PXD014414_psm_peptides.svg
plot-kde-intensity-distribution
quantmsio_cli plot plot-kde-intensity-distribution
--feature_path PXD010154-51b34353-227f-4d38-a181-6d42824de9f7.featrue.parquet
--num_samples 10
--save_path PXD014414_psm_peptides.svg
plot-bar-peptide-distribution
quantmsio_cli plot plot-bar-peptide-distribution
--feature_path PXD010154-51b34353-227f-4d38-a181-6d42824de9f7.featrue.parquet
--num_samples 10
--save_path PXD014414_psm_peptides.svg
plot-box-intensity-distribution
quantmsio_cli plot plot-box-intensity-distribution
--feature_path PXD010154-51b34353-227f-4d38-a181-6d42824de9f7.featrue.parquet
--num_samples 10
--save_path PXD014414_psm_peptides.svg